Astrometric epoch propagation#
This notebook demonstrates how to improve crossmatches between LSST DP1 objects and Gaia DR3 by accounting for stellar motion over time (epoch propagation).
Epoch propagation updates a source’s sky position from one observation epoch to another using its proper motion (and, when available, parallax and radial velocity). This should not be confused with the Equinox of the coordinate system: both catalogs use the ICRS frame with Equinox J2000.
Run this notebook on the Rubin Science Platform (or another environment where DP1 is available and LSDB is pre-installed). For DP1 access details, see: https://lsdb.io/dp1
We will:
Set up the environment and define epochs for DP1 and Gaia DR3.
Open the relevant catalogs and compute a safe search radius.
Propagate Gaia source coordinates from the Gaia epoch to the DP1 epoch.
Crossmatch DP1 objects with the propagated Gaia positions and select the best matches.
Compare against a baseline crossmatch that does not use propagation.
Visualize the difference between the two approaches.
[1]:
import lsdb
import numpy as np
from astropy import units
from astropy.coordinates import Distance, ICRS, SkyCoord
from astropy.time import Time
from dask.distributed import Client
from upath import UPath
[2]:
print(f"{lsdb.__version__ = }")
lsdb.__version__ = '0.7.3'
1. Define epochs#
We specify the observation epochs for LSST DP1 (approximately late 2024) and Gaia DR3 (2016.0). These are used to translate Gaia source positions from the Gaia epoch to the DP1 epoch.
[3]:
# DP1 was mostly October 2024, we use an approximate value here for simplicity
dp1_epoch = 2024.9
# https://gaia.aip.de/metadata/gaiadr3/
gaia_dr3_epoch = 2016.0
2. Open catalogs and gather configuration#
Open the DP1 object catalog and retrieve its matching margin. This margin helps us decide how far to search around each DP1 source when looking for Gaia counterparts. Here we use Rubin Science Platform path to DP1, please adjust as needed; see https://lsdb.io/dp1 for details.
[4]:
base_path = UPath("/rubin/lsdb_data")
dp1_object = lsdb.open_catalog(
base_path / "object_collection",
columns=[],
)
dp1_margin_arcsec = dp1_object.margin.hc_structure.catalog_info.margin_threshold
3. Derive a proper-motion filter#
Compute a maximum allowed proper motion based on the DP1 margin and the time difference between the two epochs. This helps us restrict Gaia sources to those that could plausibly still fall within the DP1 search area after propagation.
[5]:
max_pm = 1000.0 * dp1_margin_arcsec / (dp1_epoch - gaia_dr3_epoch)
max_pm
[5]:
561.797752808983
4. Load a filtered Gaia DR3 subset#
Open a Gaia DR3 catalog with the columns needed for propagation and filter out entries lacking required motion or distance information. For S3 access we use MAST’s open buckets; see https://data.lsdb.io/#Gaia/Gaia_DR3_(US-East,_S3) for details.
[6]:
gaia_dr3 = lsdb.open_catalog(
"s3://stpubdata/gaia/gaia_dr3/public/hats",
columns=["pmra", "pmdec", "radial_velocity", "parallax"],
filters=[("pm", "<", max_pm), ("parallax", ">", 0.0)],
).query("radial_velocity.notna()")
5. Define epoch propagation#
Create a helper that takes Gaia source attributes at the Gaia epoch and computes their positions at the DP1 epoch. We leverage Astropy to carry out the space‑motion propagation.
[7]:
def propagate_epoch(df, *, start, end):
if len(df) == 0:
return df.assign(ra_corr=np.array([], dtype=float), dec_corr=np.array([], dtype=float))
start_coord = SkyCoord(
ra=df["ra"].to_numpy() * units.deg,
dec=df["dec"].to_numpy() * units.deg,
pm_ra_cosdec=df["pmra"].to_numpy() * units.marcsec / units.year,
pm_dec=df["pmdec"].to_numpy() * units.marcsec / units.year,
radial_velocity=df["radial_velocity"].to_numpy() * units.km / units.s,
distance=Distance(parallax=df["parallax"].to_numpy() * units.marcsec),
obstime=Time(start, format="byear"),
frame="icrs",
)
end_coord = start_coord.apply_space_motion(new_obstime=Time(end, format="byear"))
df["ra_corr"] = end_coord.ra.deg
df["dec_corr"] = end_coord.dec.deg
return df
propagated_gaia = gaia_dr3.map_partitions(propagate_epoch, start=gaia_dr3_epoch, end=dp1_epoch)
propagated_gaia
[7]:
| pmra | pmdec | radial_velocity | parallax | ra | dec | ra_corr | dec_corr | |
|---|---|---|---|---|---|---|---|---|
| npartitions=2016 | ||||||||
| Order: 2, Pixel: 0 | double[pyarrow] | double[pyarrow] | float[pyarrow] | double[pyarrow] | double[pyarrow] | double[pyarrow] | float64 | float64 |
| Order: 2, Pixel: 1 | ... | ... | ... | ... | ... | ... | ... | ... |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
| Order: 3, Pixel: 766 | ... | ... | ... | ... | ... | ... | ... | ... |
| Order: 3, Pixel: 767 | ... | ... | ... | ... | ... | ... | ... | ... |
6. Crossmatch propagated Gaia to DP1 and select the best match#
This is a two‑step crossmatching pipeline:
We search with the largest radius that still preserves correct matches — the DP1 catalog’s margin defines this “safe” maximum. Using this margin ensures candidates that truly belong together are not missed.
We intentionally keep multiple nearby Gaia candidates per DP1 object at first. After that, we compare candidates and choose the single best one using their separation. This approach lets us propagate to the target epoch and then make an informed choice among several plausible neighbors.
[8]:
xmatch = dp1_object.crossmatch_nested(propagated_gaia, radius_arcsec=dp1_margin_arcsec, n_neighbors=20)
xmatch
[8]:
| coord_ra | coord_dec | gaia | |
|---|---|---|---|
| npartitions=389 | |||
| Order: 6, Pixel: 130 | double[pyarrow] | double[pyarrow] | nested<pmra: [double], pmdec: [double], radial... |
| Order: 8, Pixel: 2176 | ... | ... | ... |
| ... | ... | ... | ... |
| Order: 9, Pixel: 2302101 | ... | ... | ... |
| Order: 7, Pixel: 143884 | ... | ... | ... |
For each DP1 object, we then choose the closest propagated Gaia candidate within a small tolerance. This reduces multiple candidates to a single best match when possible.
[9]:
def select_best_match(dp1_ra, dp1_dec, gaia_ra, gaia_dec, *, max_dist_arcsec):
dp1_coord = SkyCoord(ra=dp1_ra * units.deg, dec=dp1_dec * units.deg)
gaia_coord = SkyCoord(ra=gaia_ra * units.deg, dec=gaia_dec * units.deg)
distance = gaia_coord.separation(dp1_coord)
idx_close = np.where(distance < max_dist_arcsec * units.arcsec)[0]
if len(idx_close) == 0:
return {"gaia_ra_corr": np.nan, "gaia_dec_corr": np.nan, "distance": np.nan}
best_idx = min(idx_close, key=distance.__getitem__)
return {
"gaia_ra_corr": gaia_ra[best_idx],
"gaia_dec_corr": gaia_dec[best_idx],
"distance": distance[best_idx].to_value(units.arcsec),
}
best_matches = (
xmatch.map_rows(
select_best_match,
columns=["coord_ra", "coord_dec", "gaia.ra_corr", "gaia.dec_corr"],
max_dist_arcsec=0.1,
append_columns=True,
row_container="args",
meta=dict.fromkeys(["gaia_ra_corr", "gaia_dec_corr", "distance"], float),
)
.map_partitions(lambda df: df.drop(columns=["gaia"]))
.query("distance.notna()")
)
best_matches
[9]:
| coord_ra | coord_dec | gaia_ra_corr | gaia_dec_corr | distance | |
|---|---|---|---|---|---|
| npartitions=389 | |||||
| Order: 6, Pixel: 130 | double[pyarrow] | double[pyarrow] | float64 | float64 | float64 |
| Order: 8, Pixel: 2176 | ... | ... | ... | ... | ... |
| ... | ... | ... | ... | ... | ... |
| Order: 9, Pixel: 2302101 | ... | ... | ... | ... | ... |
| Order: 7, Pixel: 143884 | ... | ... | ... | ... | ... |
7. Baseline: crossmatch without propagation#
As a point of comparison, perform a standard positional crossmatch using the Gaia positions at their original epoch (no propagation).
[10]:
naive_matches = dp1_object.crossmatch(gaia_dr3, radius_arcsec=0.1, suffix_method="overlapping_columns")
8. Compute results with a local Dask client#
All steps above are lazily evaluated. Here we actually run both pipelines — with propagation and without — and collect the results into Pandas DataFrames. We also print the number of matches from each pipeline.
[11]:
with Client(n_workers=4, threads_per_worker=1) as client:
display(client)
best_matches_df = best_matches.compute()
print(f"Propagated matches: {len(best_matches_df)}")
naive_matches_df = naive_matches.compute()
print(f"Non-propagated matches: {len(naive_matches_df)}")
distributed.scheduler INFO: State start
distributed.scheduler INFO: Scheduler at: tcp://127.0.0.1:46535
distributed.scheduler INFO: dashboard at: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/8787/status
distributed.scheduler INFO: Registering Worker plugin shuffle
distributed.nanny INFO: Start Nanny at: 'tcp://127.0.0.1:37929'
distributed.nanny INFO: Start Nanny at: 'tcp://127.0.0.1:32893'
distributed.nanny INFO: Start Nanny at: 'tcp://127.0.0.1:39689'
distributed.nanny INFO: Start Nanny at: 'tcp://127.0.0.1:37729'
distributed.scheduler INFO: Register worker addr: tcp://127.0.0.1:44929 name: 0
distributed.scheduler INFO: Starting worker compute stream, tcp://127.0.0.1:44929
distributed.core INFO: Starting established connection to tcp://127.0.0.1:52778
distributed.scheduler INFO: Register worker addr: tcp://127.0.0.1:46017 name: 1
distributed.scheduler INFO: Starting worker compute stream, tcp://127.0.0.1:46017
distributed.core INFO: Starting established connection to tcp://127.0.0.1:52782
distributed.scheduler INFO: Register worker addr: tcp://127.0.0.1:35485 name: 2
distributed.scheduler INFO: Starting worker compute stream, tcp://127.0.0.1:35485
distributed.core INFO: Starting established connection to tcp://127.0.0.1:52792
distributed.scheduler INFO: Register worker addr: tcp://127.0.0.1:44707 name: 3
distributed.scheduler INFO: Starting worker compute stream, tcp://127.0.0.1:44707
distributed.core INFO: Starting established connection to tcp://127.0.0.1:52806
distributed.scheduler INFO: Receive client connection: Client-08256a2e-da07-11f0-827e-af10543ff312
distributed.core INFO: Starting established connection to tcp://127.0.0.1:52808
Client
Client-08256a2e-da07-11f0-827e-af10543ff312
| Connection method: Cluster object | Cluster type: distributed.LocalCluster |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/8787/status |
Cluster Info
LocalCluster
15ba409d
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/8787/status | Workers: 4 |
| Total threads: 4 | Total memory: 16.00 GiB |
| Status: running | Using processes: True |
Scheduler Info
Scheduler
Scheduler-d9cd6483-becd-4cd3-b6df-1656560ab186
| Comm: tcp://127.0.0.1:46535 | Workers: 0 |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/8787/status | Total threads: 0 |
| Started: Just now | Total memory: 0 B |
Workers
Worker: 0
| Comm: tcp://127.0.0.1:44929 | Total threads: 1 |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/41041/status | Memory: 4.00 GiB |
| Nanny: tcp://127.0.0.1:37929 | |
| Local directory: /tmp/dask-scratch-space/worker-5jwwnhta | |
Worker: 1
| Comm: tcp://127.0.0.1:46017 | Total threads: 1 |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/35583/status | Memory: 4.00 GiB |
| Nanny: tcp://127.0.0.1:32893 | |
| Local directory: /tmp/dask-scratch-space/worker-9xafhnm1 | |
Worker: 2
| Comm: tcp://127.0.0.1:35485 | Total threads: 1 |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/45399/status | Memory: 4.00 GiB |
| Nanny: tcp://127.0.0.1:39689 | |
| Local directory: /tmp/dask-scratch-space/worker-ijjn4t4_ | |
Worker: 3
| Comm: tcp://127.0.0.1:44707 | Total threads: 1 |
| Dashboard: https://malanchev.nb.data.lsst.cloud/nb/user/malanchev/proxy/46427/status | Memory: 4.00 GiB |
| Nanny: tcp://127.0.0.1:37729 | |
| Local directory: /tmp/dask-scratch-space/worker-nl9b83jc | |
Propagated matches: 1492
Non-propagated matches: 957
distributed.scheduler INFO: Remove client Client-08256a2e-da07-11f0-827e-af10543ff312
distributed.core INFO: Received 'close-stream' from tcp://127.0.0.1:52808; closing.
distributed.scheduler INFO: Remove client Client-08256a2e-da07-11f0-827e-af10543ff312
distributed.scheduler INFO: Close client connection: Client-08256a2e-da07-11f0-827e-af10543ff312
distributed.scheduler INFO: Retire worker addresses (stimulus_id='retire-workers-1765838977.415765') (0, 1, 2, 3)
distributed.nanny INFO: Closing Nanny at 'tcp://127.0.0.1:37929'. Reason: nanny-close
distributed.nanny INFO: Nanny asking worker to close. Reason: nanny-close
distributed.nanny INFO: Closing Nanny at 'tcp://127.0.0.1:32893'. Reason: nanny-close
distributed.nanny INFO: Nanny asking worker to close. Reason: nanny-close
distributed.nanny INFO: Closing Nanny at 'tcp://127.0.0.1:39689'. Reason: nanny-close
distributed.nanny INFO: Nanny asking worker to close. Reason: nanny-close
distributed.nanny INFO: Closing Nanny at 'tcp://127.0.0.1:37729'. Reason: nanny-close
distributed.nanny INFO: Nanny asking worker to close. Reason: nanny-close
distributed.core INFO: Received 'close-stream' from tcp://127.0.0.1:52782; closing.
distributed.core INFO: Received 'close-stream' from tcp://127.0.0.1:52792; closing.
distributed.scheduler INFO: Remove worker addr: tcp://127.0.0.1:46017 name: 1 (stimulus_id='handle-worker-cleanup-1765838977.4288983')
distributed.scheduler INFO: Remove worker addr: tcp://127.0.0.1:35485 name: 2 (stimulus_id='handle-worker-cleanup-1765838977.4302118')
distributed.core INFO: Received 'close-stream' from tcp://127.0.0.1:52806; closing.
distributed.scheduler INFO: Remove worker addr: tcp://127.0.0.1:44707 name: 3 (stimulus_id='handle-worker-cleanup-1765838977.4333227')
distributed.core INFO: Received 'close-stream' from tcp://127.0.0.1:52778; closing.
distributed.scheduler INFO: Remove worker addr: tcp://127.0.0.1:44929 name: 0 (stimulus_id='handle-worker-cleanup-1765838977.4580626')
distributed.scheduler INFO: Lost all workers
distributed.nanny INFO: Nanny at 'tcp://127.0.0.1:37729' closed.
distributed.nanny INFO: Nanny at 'tcp://127.0.0.1:37929' closed.
distributed.nanny INFO: Nanny at 'tcp://127.0.0.1:32893' closed.
distributed.nanny INFO: Nanny at 'tcp://127.0.0.1:39689' closed.
distributed.scheduler INFO: Closing scheduler. Reason: unknown
distributed.scheduler INFO: Scheduler closing all comms
9. Plot separation distribution#
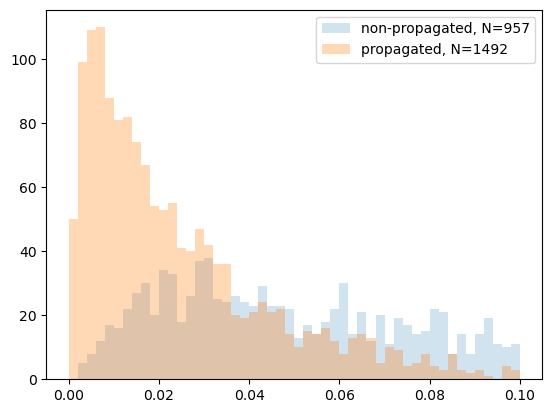
Plot histograms of match separations for both approaches and highlight the effect of propagation. In this run, the propagated pipeline yields more matches (1492 vs 957), and the separation distribution is tighter (more matches at smaller separations).
[12]:
import matplotlib.pyplot as plt
plt.hist(
naive_matches_df["_dist_arcsec"],
bins=np.linspace(0, 0.1, 51),
histtype="stepfilled",
alpha=0.2,
label=f"non-propagated, N={len(naive_matches_df)}",
)
plt.hist(
best_matches_df["distance"],
bins=np.linspace(0, 0.1, 51),
histtype="stepfilled",
alpha=0.3,
label=f"propagated, N={len(best_matches_df)}",
)
plt.legend()
[12]:
<matplotlib.legend.Legend at 0x7f78a49b41a0>
[ ]:
