Joining catalogs#
In this tutorial, we will:
demonstrate how to perform a JOIN on two catalogs
Note that this is different from a crossmatch, because the two catalogs share unique identifiers and we can match those values directly. This will still use the spatial properties to perform the JOIN on a per-partition basis.
Introduction#
Gaia is a space telescope that provides excellent astrometric precision, and so is used for determining parallax distances to nearby stars. The parallax distances are not available in all data products, however.
In this notebook, we join Gaia with Gaia Early Data Release 3 (EDR3) and compute the ratio between the distances given by their parallax and r_med_geo columns, respectively.
[1]:
import lsdb
from lsdb import ConeSearch
1. Load the catalogs#
First we load Gaia with its objects source_id, their positions and parallax columns.
[ ]:
gaia = lsdb.open_catalog(
"https://data.lsdb.io/hats/gaia_dr3",
columns=["source_id", "ra", "dec", "parallax"],
search_filter=ConeSearch(ra=0, dec=0, radius_arcsec=10 * 3600),
)
gaia
| source_id | ra | dec | parallax | |
|---|---|---|---|---|
| npartitions=4 | ||||
| Order: 2, Pixel: 67 | int64[pyarrow] | double[pyarrow] | double[pyarrow] | double[pyarrow] |
| Order: 2, Pixel: 70 | ... | ... | ... | ... |
| Order: 2, Pixel: 73 | ... | ... | ... | ... |
| Order: 2, Pixel: 76 | ... | ... | ... | ... |
We will do the same with Gaia EDR3 but the distance column we will use is called r_med_geo, the median of the geometric distance estimate.
[ ]:
gaia_edr3 = lsdb.open_catalog(
"https://data.lsdb.io/hats/gaia_edr3_distances",
columns=["source_id", "ra", "dec", "r_med_geo"],
search_filter=ConeSearch(ra=0, dec=0, radius_arcsec=10 * 3600),
)
gaia_edr3
| source_id | ra | dec | r_med_geo | |
|---|---|---|---|---|
| npartitions=4 | ||||
| Order: 2, Pixel: 67 | int64[pyarrow] | double[pyarrow] | double[pyarrow] | double[pyarrow] |
| Order: 2, Pixel: 70 | ... | ... | ... | ... |
| Order: 2, Pixel: 73 | ... | ... | ... | ... |
| Order: 2, Pixel: 76 | ... | ... | ... | ... |
2. Join Operation#
We are now able to join both catalogs on the source_id column, as follows:
[4]:
joined = gaia.join(gaia_edr3, left_on="source_id", right_on="source_id")
joined
/astro/users/olynn/lsdb/src/lsdb/catalog/catalog.py:840: FutureWarning: The default suffix behavior will change from applying suffixes to all columns to only applying suffixes to overlapping columns in a future release.To maintain the current behavior, explicitly set `suffix_method='all_columns'`. To change to the new behavior, set `suffix_method='overlapping_columns'`.
warnings.warn(
[4]:
| source_id_gaia | ra_gaia | dec_gaia | parallax_gaia | source_id_gaia_edr3_distances | ra_gaia_edr3_distances | dec_gaia_edr3_distances | r_med_geo_gaia_edr3_distances | |
|---|---|---|---|---|---|---|---|---|
| npartitions=4 | ||||||||
| Order: 2, Pixel: 67 | int64[pyarrow] | double[pyarrow] | double[pyarrow] | double[pyarrow] | int64[pyarrow] | double[pyarrow] | double[pyarrow] | double[pyarrow] |
| Order: 2, Pixel: 70 | ... | ... | ... | ... | ... | ... | ... | ... |
| Order: 2, Pixel: 73 | ... | ... | ... | ... | ... | ... | ... | ... |
| Order: 2, Pixel: 76 | ... | ... | ... | ... | ... | ... | ... | ... |
3. Joint analysis#
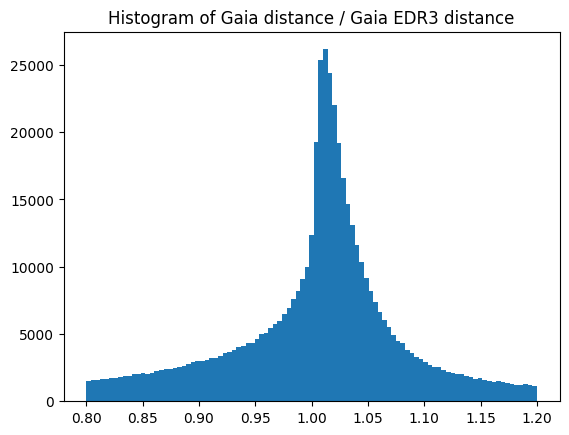
Let’s calculate a histogram with the ratio in catalog distances.
[5]:
results = (1e3 / joined["parallax_gaia"]) / joined["r_med_geo_gaia_edr3_distances"]
ratios = results.compute().to_numpy()
ratios
[5]:
array([ 1.04369179, 0.99821673, 0.96884423, ..., 1.03218566,
147.42381857, 1.41197562], shape=(825678,))
[6]:
import numpy as np
import matplotlib.pyplot as plt
plt.hist(ratios, bins=np.linspace(0.8, 1.2, 100))
plt.title("Histogram of Gaia distance / Gaia EDR3 distance")
plt.show()
About#
Authors: Sandro Campos
Last updated on: Oct 27, 2025
If you use lsdb for published research, please cite following instructions.
