Using Index Tables#
In this tutorial, we will:
discuss the purpose and scope of secondary index tables in LSDB
explore using index tables to find GAIA objects, by designation
Introduction#
HATS catalogs are partitioned spatially, typically on right ascension and declination. This makes finding objects in a particular area of the sky very straightforward. However, you may occassionally only know the object by the survey-assigned identifier.
HATS supports creating additional secondary index tables, stored separately from the HATS catalog. You can find more information about creating these index tables in the hats-import documentation, or request that your archive provider create an index table for your use.
1. Load a catalog collection#
HATS catalogs and supplemental tables can be gathered together into a “catalog collection”. In this way, margins and index tables can be accessed without needing to know their subdirectories.
GAIA is available on data.lsdb.io in the form of a catalog collection.
[5]:
import lsdb
gaia = lsdb.open_catalog("https://data.lsdb.io/hats/gaia_dr3/")
We can inspect the catalog’s collection properties and find which columns have secondary index tables.
[6]:
gaia.hc_collection.all_indexes.keys()
[6]:
dict_keys(['source_id'])
2. Find items by ID#
Let’s create some list of designations that we’re interested in. The ones here have no special signficance, but are not near each other on the sphere.
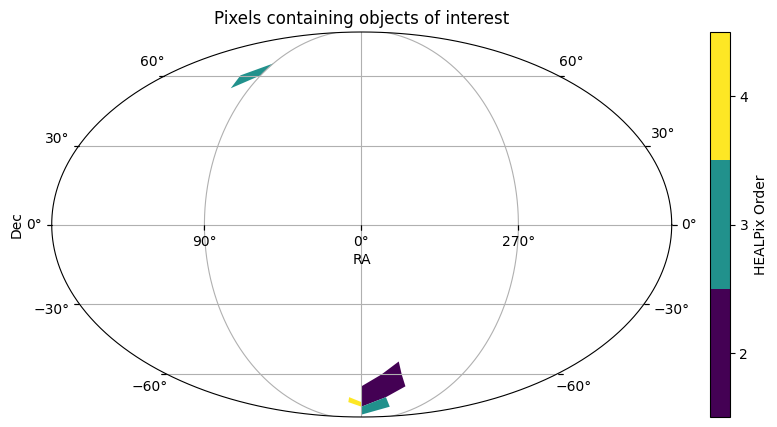
This small set of GAIA designations are in just a handful of partitions. Let’s check the index catalog, and see if we can reduce this list of IDs into just those pixels of interest.
[ ]:
ids = [
6350084614282952448,
4684296460655809408,
4684296460655681280,
6350084408124522368,
6379030430412267264,
6397962092201358080,
999999988604363776,
]
[8]:
id_filtered = gaia.id_search(values={"source_id": ids})
Searching for source_id=[6350084614282952448, 4684296460655809408, 4684296460655681280, 6350084408124522368, 6379030430412267264, 6397962092201358080, 999999988604363776]
[9]:
id_filtered.plot_pixels(plot_title="Pixels containing objects of interest")
[9]:
(<Figure size 1000x500 with 2 Axes>,
<WCSAxes: title={'center': 'Pixels containing objects of interest'}>)
[10]:
id_filtered.compute()
[10]:
| solution_id | designation | source_id | random_index | ref_epoch | ra | ra_error | dec | dec_error | parallax | parallax_error | parallax_over_error | pm | pmra | pmra_error | pmdec | pmdec_error | ra_dec_corr | ra_parallax_corr | ra_pmra_corr | ra_pmdec_corr | dec_parallax_corr | dec_pmra_corr | dec_pmdec_corr | parallax_pmra_corr | parallax_pmdec_corr | pmra_pmdec_corr | astrometric_n_obs_al | astrometric_n_obs_ac | astrometric_n_good_obs_al | astrometric_n_bad_obs_al | astrometric_gof_al | astrometric_chi2_al | astrometric_excess_noise | astrometric_excess_noise_sig | astrometric_params_solved | astrometric_primary_flag | nu_eff_used_in_astrometry | pseudocolour | pseudocolour_error | ra_pseudocolour_corr | dec_pseudocolour_corr | parallax_pseudocolour_corr | pmra_pseudocolour_corr | pmdec_pseudocolour_corr | astrometric_matched_transits | visibility_periods_used | astrometric_sigma5d_max | matched_transits | new_matched_transits | matched_transits_removed | ipd_gof_harmonic_amplitude | ipd_gof_harmonic_phase | ipd_frac_multi_peak | ipd_frac_odd_win | ruwe | scan_direction_strength_k1 | scan_direction_strength_k2 | scan_direction_strength_k3 | scan_direction_strength_k4 | scan_direction_mean_k1 | scan_direction_mean_k2 | scan_direction_mean_k3 | scan_direction_mean_k4 | duplicated_source | phot_g_n_obs | phot_g_mean_flux | phot_g_mean_flux_error | phot_g_mean_flux_over_error | phot_g_mean_mag | phot_bp_n_obs | phot_bp_mean_flux | phot_bp_mean_flux_error | phot_bp_mean_flux_over_error | phot_bp_mean_mag | phot_rp_n_obs | phot_rp_mean_flux | phot_rp_mean_flux_error | phot_rp_mean_flux_over_error | phot_rp_mean_mag | phot_bp_rp_excess_factor | phot_bp_n_contaminated_transits | phot_bp_n_blended_transits | phot_rp_n_contaminated_transits | phot_rp_n_blended_transits | phot_proc_mode | bp_rp | bp_g | g_rp | radial_velocity | radial_velocity_error | rv_method_used | rv_nb_transits | rv_nb_deblended_transits | rv_visibility_periods_used | rv_expected_sig_to_noise | rv_renormalised_gof | rv_chisq_pvalue | rv_time_duration | rv_amplitude_robust | rv_template_teff | rv_template_logg | rv_template_fe_h | rv_atm_param_origin | vbroad | vbroad_error | vbroad_nb_transits | grvs_mag | grvs_mag_error | grvs_mag_nb_transits | rvs_spec_sig_to_noise | phot_variable_flag | l | b | ecl_lon | ecl_lat | in_qso_candidates | in_galaxy_candidates | non_single_star | has_xp_continuous | has_xp_sampled | has_rvs | has_epoch_photometry | has_epoch_rv | has_mcmc_gspphot | has_mcmc_msc | in_andromeda_survey | classprob_dsc_combmod_quasar | classprob_dsc_combmod_galaxy | classprob_dsc_combmod_star | teff_gspphot | teff_gspphot_lower | teff_gspphot_upper | logg_gspphot | logg_gspphot_lower | logg_gspphot_upper | mh_gspphot | mh_gspphot_lower | mh_gspphot_upper | distance_gspphot | distance_gspphot_lower | distance_gspphot_upper | azero_gspphot | azero_gspphot_lower | azero_gspphot_upper | ag_gspphot | ag_gspphot_lower | ag_gspphot_upper | ebpminrp_gspphot | ebpminrp_gspphot_lower | ebpminrp_gspphot_upper | libname_gspphot | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| _healpix_29 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 500000002326108747 | 1636148068921376768 | Gaia DR3 999999988604363776 | 999999988604363776 | 1146519553 | 2016.0 | 104.910499 | 0.416448 | 55.969686 | 0.392039 | 1.501113 | 0.662432 | 2.266063 | 1.077911 | 1.004717 | 0.509705 | -0.39043 | 0.407673 | 0.166491 | -0.283951 | -0.210769 | -0.166695 | -0.395157 | -0.383997 | -0.082387 | 0.571293 | -0.097417 | -0.023115 | 279 | 0 | 278 | 1 | 0.190794 | 302.275909 | 0.918131 | 0.573547 | 95 | False | <NA> | 1.231943 | 0.12604 | 0.02382 | -0.028157 | -0.177176 | -0.060274 | 0.381471 | 32 | 19 | 0.800948 | 34 | 34 | 0 | 0.043367 | 8.364323 | 0 | 0 | 1.006963 | 0.065047 | 0.08709 | 0.127981 | 0.628179 | -134.500015 | 20.566761 | 23.053226 | -39.491501 | False | 284 | 156.458699 | 0.820939 | 190.584991 | 20.201368 | 26 | 56.254621 | 9.234386 | 6.091863 | 20.963146 | 28 | 168.634235 | 9.440611 | 17.86264 | 19.180531 | 1.437369 | 0 | 0 | 0 | 0 | 0 | 1.782616 | 0.761778 | 1.020838 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 160.447923 | 23.310772 | 99.891286 | 33.041242 | False | False | 0 | False | False | False | False | False | False | False | False | 0.0 | 0.0 | 0.999994 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> |
| 2342148232994970763 | 1636148068921376768 | Gaia DR3 4684296460655681280 | 4684296460655681280 | 935088391 | 2016.0 | 7.756788 | 0.075319 | -75.621828 | 0.069098 | 0.249487 | 0.081103 | 3.076187 | 5.066541 | 4.582298 | 0.096344 | 2.16157 | 0.086174 | -0.052206 | 0.15723 | -0.084423 | 0.031885 | -0.040616 | 0.067726 | 0.007037 | -0.097317 | -0.099672 | 0.148652 | 339 | 0 | 338 | 1 | 0.858621 | 369.676392 | 0.0 | 0.0 | 31 | False | 1.541551 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 39 | 27 | 0.137921 | 43 | 15 | 0 | 0.00746 | 77.971634 | 0 | 0 | 1.032443 | 0.09834 | 0.118443 | 0.059276 | 0.042126 | -6.52272 | 28.822557 | -7.548758 | -35.488499 | False | 354 | 1425.008016 | 1.483123 | 960.815674 | 17.802824 | 35 | 720.948503 | 8.592239 | 83.906937 | 18.193781 | 36 | 974.007199 | 10.457192 | 93.142326 | 17.276489 | 1.189436 | 0 | 1 | 0 | 0 | 0 | 0.917292 | 0.390957 | 0.526335 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 304.62018 | -41.439446 | 304.758315 | -64.432345 | False | False | 0 | False | False | False | False | False | True | True | False | 0.0 | 0.0 | 0.999989 | 5312.479004 | 5298.666992 | 5326.42334 | 4.701 | 4.6734 | 4.7276 | -0.9948 | -1.1187 | -0.9004 | 2123.999512 | 2045.833008 | 2180.535645 | 0.006 | 0.0012 | 0.0158 | 0.005 | 0.001 | 0.013 | 0.0027 | 0.0005 | 0.0071 | MARCS |
| 2342148241066740563 | 1636148068921376768 | Gaia DR3 4684296460655809408 | 4684296460655809408 | 930596526 | 2016.0 | 7.74122 | 3.736654 | -75.619753 | 3.646013 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | -0.2702 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 175 | 0 | 175 | 0 | 31.654589 | 1673.466064 | 30.255457 | 72.205276 | 3 | False | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 20 | 18 | 7.92256 | 21 | 8 | 1 | 0.390671 | 52.280476 | 0 | 0 | <NA> | 0.153033 | 0.211406 | 0.028252 | 0.119918 | 65.863274 | 44.792297 | 56.646534 | 10.881159 | False | 180 | 84.509249 | 1.814717 | 46.568825 | 20.870106 | 17 | 57.310949 | 7.826055 | 7.323096 | 20.942947 | 16 | 62.458944 | 11.260726 | 5.546618 | 20.258909 | 1.41724 | 0 | 0 | 0 | 1 | 0 | 0.684038 | 0.072842 | 0.611197 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 304.625601 | -41.4411 | 304.759053 | -64.427969 | False | False | 0 | False | False | False | False | False | False | False | False | 0.017581 | 0.000203 | 0.979297 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> |
| 3175042221187998424 | 1636148068921376768 | Gaia DR3 6350084408124522368 | 6350084408124522368 | 64296681 | 2016.0 | 359.838284 | 0.249825 | -83.92197 | 0.239047 | 0.656007 | 0.264131 | 2.483639 | 8.980813 | 8.351725 | 0.293432 | -3.302074 | 0.362815 | 0.019055 | 0.221716 | -0.445573 | -0.099703 | -0.217621 | -0.027286 | -0.071756 | -0.271884 | -0.025407 | -0.070919 | 301 | 0 | 300 | 1 | 0.585375 | 334.195038 | 0.147555 | 0.036804 | 95 | False | <NA> | 1.259485 | 0.073974 | -0.156117 | 0.116803 | -0.035608 | 0.215098 | -0.012293 | 35 | 23 | 0.50419 | 42 | 16 | 0 | 0.022326 | 134.932983 | 0 | 0 | 1.023095 | 0.017062 | 0.021049 | 0.152066 | 0.05543 | 164.156006 | 50.43185 | -54.592541 | -0.473558 | False | 345 | 296.790726 | 0.879197 | 337.570221 | 19.506241 | 37 | 104.582573 | 7.116625 | 14.69553 | 20.289894 | 36 | 292.777679 | 8.608073 | 34.011986 | 18.581551 | 1.338857 | 0 | 1 | 0 | 0 | 0 | 1.708344 | 0.783653 | 0.92469 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 304.56271 | -33.04051 | 284.976231 | -65.811972 | False | False | 0 | False | False | False | False | False | False | False | False | 0.0 | 0.0 | 0.999984 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> |
| 3175042313799520523 | 1636148068921376768 | Gaia DR3 6350084614282952448 | 6350084614282952448 | 1750910513 | 2016.0 | 359.435728 | 0.510138 | -83.921882 | 0.492162 | -0.272154 | 0.53593 | -0.507817 | 10.734181 | 9.925715 | 0.625717 | -4.086909 | 0.660092 | 0.201444 | 0.051819 | -0.209568 | -0.009297 | -0.123529 | -0.019864 | -0.0042 | -0.226232 | 0.009305 | 0.178916 | 256 | 0 | 254 | 2 | -1.012652 | 240.363922 | 0.0 | 0.0 | 95 | False | <NA> | 1.433166 | 0.169743 | -0.19527 | 0.127554 | -0.119694 | 0.130828 | -0.253519 | 30 | 22 | 0.972411 | 35 | 12 | 0 | 0.064661 | 83.889114 | 0 | 0 | 0.953554 | 0.168301 | 0.119298 | 0.125173 | 0.172718 | -11.14678 | -31.648802 | 52.093277 | -34.022755 | False | 294 | 122.977285 | 0.759025 | 162.020157 | 20.462805 | 27 | 51.462335 | 6.843296 | 7.52011 | 21.059818 | 32 | 105.089089 | 6.625872 | 15.860417 | 19.694002 | 1.273011 | 0 | 1 | 0 | 0 | 0 | 1.365816 | 0.597013 | 0.768803 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 304.612058 | -33.030245 | 284.951178 | -65.770605 | False | False | 0 | False | False | False | False | False | False | False | False | 0.0 | 0.0 | 0.999994 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> |
| 3189515220382416186 | 1636148068921376768 | Gaia DR3 6379030430412267264 | 6379030430412267264 | 1257926240 | 2016.0 | 348.261377 | 0.101749 | -73.622147 | 0.093109 | 0.199036 | 0.11661 | 1.706857 | 12.912383 | 0.788176 | 0.139862 | -12.888305 | 0.131521 | -0.091888 | 0.040155 | -0.378473 | 0.129075 | -0.192607 | 0.177191 | -0.033713 | -0.255931 | -0.274566 | -0.03595 | 356 | 0 | 354 | 2 | 2.310223 | 424.929199 | 0.4559 | 2.129916 | 95 | False | <NA> | 1.508331 | 0.028194 | -0.056774 | 0.047647 | -0.050193 | 0.265732 | -0.009968 | 41 | 25 | 0.201031 | 49 | 49 | 0 | 0.011486 | 97.635696 | 0 | 0 | 1.087849 | 0.022523 | 0.068986 | 0.154577 | 0.181862 | 115.995918 | 34.802052 | -32.362061 | -33.449848 | False | 403 | 1055.450301 | 1.218892 | 865.909363 | 18.128773 | 42 | 560.927295 | 7.779328 | 72.104851 | 18.466276 | 43 | 721.210458 | 9.912146 | 72.760277 | 17.602741 | 1.214778 | 0 | 3 | 0 | 2 | 0 | 0.863535 | 0.337503 | 0.526031 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 311.980909 | -41.732829 | 302.444948 | -59.029747 | False | False | 0 | False | False | False | False | False | True | True | False | 0.0 | 0.0 | 0.99997 | 5460.65918 | 5448.008301 | 5475.993652 | 4.6978 | 4.6729 | 4.7202 | -1.179 | -1.3243 | -1.0133 | 2611.122314 | 2536.43457 | 2699.760254 | 0.0061 | 0.0015 | 0.0155 | 0.005 | 0.0012 | 0.0129 | 0.0027 | 0.0006 | 0.007 | MARCS |
| 3192976252233877374 | 1636148068921376768 | Gaia DR3 6397962092201358080 | 6397962092201358080 | 150735067 | 2016.0 | 336.476393 | 0.079348 | -67.482415 | 0.09224 | 0.239489 | 0.106303 | 2.252891 | 9.269107 | 3.705983 | 0.102108 | -8.496001 | 0.118224 | -0.347174 | 0.171825 | -0.240193 | 0.223929 | -0.209054 | 0.258152 | -0.222417 | -0.103846 | -0.083716 | -0.355912 | 429 | 0 | 428 | 1 | 0.089336 | 443.150299 | 0.0 | 0.0 | 31 | False | 1.497898 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 49 | 30 | 0.183705 | 52 | 19 | 0 | 0.027644 | 135.447586 | 0 | 0 | 1.002284 | 0.020995 | 0.255316 | 0.075802 | 0.211137 | -6.713617 | 52.50206 | -28.069695 | -26.521244 | False | 441 | 1036.302226 | 1.158028 | 894.885193 | 18.148651 | 48 | 485.30595 | 10.56437 | 45.937992 | 18.623503 | 47 | 765.113141 | 9.96217 | 76.801857 | 17.538582 | 1.206616 | 0 | 2 | 0 | 3 | 0 | 1.084921 | 0.474852 | 0.610069 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | NOT_AVAILABLE | 321.365755 | -44.076405 | 304.669415 | -51.88134 | False | False | 0 | False | False | False | False | False | True | True | False | 0.0 | 0.0 | 0.99999 | 5004.919434 | 4993.008789 | 5016.689941 | 4.6809 | 4.6709 | 4.6911 | -0.3503 | -0.4262 | -0.2778 | 2311.165527 | 2246.936768 | 2370.423584 | 0.0026 | 0.0005 | 0.0091 | 0.0021 | 0.0004 | 0.0073 | 0.0011 | 0.0002 | 0.0039 | MARCS |
7 rows × 152 columns
About#
Authors: Melissa DeLucchi
Last updated /verified on: October 27, 2025
If you use lsdb for published research, please cite following instructions.
